How To Make Guide Rna For Crispr involves precise design and execution, crucial for successful genome editing. At CONDUCT.EDU.VN, we provide comprehensive resources to navigate this process, ensuring accuracy and minimizing off-target effects, thus enhancing your research outcomes and fostering ethical scientific practices. Learn about gRNA design, synthesis, and validation to achieve optimal results in your CRISPR experiments and enhance your understanding of genome editing principles.
1. Understanding CRISPR and Guide RNA (gRNA)
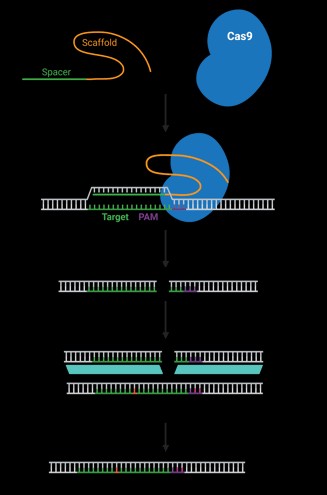
The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) system has revolutionized gene editing, offering unprecedented precision and efficiency in modifying DNA sequences. At the heart of this technology lies the guide RNA (gRNA), a short RNA sequence that directs the Cas9 enzyme to a specific location in the genome. Understanding the role of gRNA and its design principles is crucial for successful CRISPR experiments.
1.1. The Role of gRNA in CRISPR-Cas9 Systems
The CRISPR-Cas9 system functions as a molecular scalpel, enabling researchers to target and modify specific DNA sequences within a cell. The gRNA acts as a guide, directing the Cas9 enzyme to the desired location on the genome. This precision is achieved through the gRNA’s sequence complementarity to the target DNA sequence. Once the gRNA binds to the target site, the Cas9 enzyme cleaves the DNA, allowing for gene editing or modification.
1.2. Essential Components of gRNA
A gRNA typically consists of two essential components:
-
CRISPR RNA (crRNA): This is a short RNA sequence, typically around 20 nucleotides long, that is complementary to the target DNA sequence. The crRNA determines the specificity of the CRISPR-Cas9 system by guiding the Cas9 enzyme to the intended genomic location.
-
Trans-activating crRNA (tracrRNA): This longer RNA sequence acts as a scaffold, binding to the Cas9 enzyme and facilitating the formation of a functional complex. The tracrRNA is essential for the stability and activity of the CRISPR-Cas9 system.
The fusion of crRNA and tracrRNA results in a single guide RNA (sgRNA), streamlining the design and delivery process. The sgRNA simplifies the process of guiding Cas9, making it easier to use in experiments.
1.3. Importance of gRNA Design
Effective gRNA design is paramount for successful CRISPR experiments. A well-designed gRNA ensures high on-target activity, minimizing off-target effects and maximizing the efficiency of gene editing. Several factors must be considered during gRNA design, including target specificity, efficacy, and potential off-target sites. By carefully selecting and optimizing gRNA sequences, researchers can enhance the precision and reliability of CRISPR-Cas9-mediated genome editing.
2. Designing Your gRNA: Key Considerations
Designing an effective gRNA for CRISPR-Cas9 experiments requires careful consideration of several factors. These include selecting the target sequence, minimizing off-target effects, and optimizing gRNA efficacy. The following guidelines will help you create gRNAs that enhance the precision and success of your gene editing projects.
2.1. Selecting the Target Sequence
The first step in designing a gRNA is selecting the target sequence, which is a 20-nucleotide sequence that is complementary to the DNA region you want to edit. Here are some essential considerations:
-
Location within the Gene: The location of the target sequence within the gene is crucial. For gene knockout experiments, target sites near the start codon are preferred to maximize the disruption of protein translation. Avoid targeting sites close to the 3′ end of the gene to prevent the production of truncated but functional proteins. As noted by Doench et al. (2017), restricting gRNAs to 5 – 65% of the protein-coding region will still provide many options.
-
Proximity to PAM Sequence: The target sequence must be adjacent to a Protospacer Adjacent Motif (PAM) sequence. For Streptococcus pyogenes Cas9 (SpCas9), the PAM sequence is NGG (where N can be any nucleotide). The Cas9 enzyme recognizes and binds to the PAM sequence, which is essential for DNA cleavage.
-
Strand Selection: SpCas9 can target both the coding and non-coding strands of DNA with equal efficacy. Choose the strand that yields the best target sequence with minimal off-target potential.
2.2. Minimizing Off-Target Effects
Off-target effects occur when the gRNA binds to DNA sequences similar to the target sequence, leading to unintended edits at other locations in the genome. Minimizing these effects is critical for the accuracy and reliability of CRISPR experiments.
-
Sequence Specificity: Select target sequences with minimal homology to other regions of the genome. Use bioinformatics tools to scan the genome for potential off-target sites. These tools can identify sequences with one, two, or three mismatches compared to the target sequence.
-
Mismatch Tolerance: Different positions within the gRNA sequence have varying tolerance for mismatches. Mismatches near the PAM sequence are more likely to prevent Cas9 binding and cleavage, while mismatches at the distal end of the gRNA are less disruptive.
-
Modified Cas9 Enzymes: Consider using high-fidelity Cas9 variants that have been engineered to reduce off-target activity. These variants have altered amino acid sequences that increase their specificity for the target DNA.
2.3. Optimizing gRNA Efficacy
The efficacy of a gRNA refers to its ability to efficiently guide Cas9 to the target site and induce DNA cleavage. Several factors can influence gRNA efficacy, including:
-
GC Content: gRNAs with a GC content between 30% and 70% tend to be more effective. High GC content can lead to secondary structures that hinder gRNA binding, while low GC content can reduce the stability of the gRNA-DNA complex.
-
Secondary Structures: Avoid gRNA sequences that are predicted to form strong secondary structures, such as hairpins or stem-loops. These structures can interfere with gRNA binding to the target DNA.
-
Position-Specific Preferences: Certain nucleotides at specific positions within the gRNA sequence can enhance or reduce efficacy. For example, a guanine (G) at the 5′ end of the gRNA can improve its stability and activity.
2.4. Using Online gRNA Design Tools
Several online tools are available to assist in gRNA design. These tools incorporate algorithms and databases to predict on-target activity and potential off-target sites. Some popular gRNA design tools include:
-
CHOPCHOP: This tool allows you to input a gene sequence and identifies potential gRNA target sites, considering factors such as GC content and off-target potential.
-
CRISPR Design Tool (MIT): Developed by the Zhang lab at MIT, this tool provides comprehensive gRNA design and off-target analysis.
-
Benchling: Benchling is a cloud-based platform that offers a suite of tools for designing and managing CRISPR experiments, including gRNA design.
By leveraging these tools, researchers can streamline the gRNA design process and increase the likelihood of success in their CRISPR experiments. At CONDUCT.EDU.VN, we emphasize the importance of using reliable and validated resources to ensure the accuracy and ethical conduct of scientific research.
3. Methods for gRNA Synthesis
After designing your gRNA, the next step is to synthesize it. Several methods are available for gRNA synthesis, each with its own advantages and disadvantages. Here are the most common approaches:
3.1. In Vitro Transcription
In vitro transcription is a popular method for generating gRNAs. This method involves using a DNA template containing a promoter sequence recognized by an RNA polymerase, such as T7 or SP6. The RNA polymerase transcribes the DNA template into RNA, producing the gRNA.
Procedure:
-
Prepare the DNA Template: The DNA template should include the promoter sequence, the gRNA sequence, and a terminator sequence. This template can be generated by PCR or obtained from a plasmid.
-
Perform In Vitro Transcription: Use a commercially available in vitro transcription kit, following the manufacturer’s instructions. Typically, the reaction mixture includes the DNA template, RNA polymerase, NTPs (nucleotide triphosphates), and a reaction buffer.
-
DNase Treatment: After transcription, treat the reaction with DNase I to remove the DNA template.
-
RNA Purification: Purify the RNA using a column-based purification kit or phenol-chloroform extraction.
-
Quality Control: Assess the quality and concentration of the synthesized gRNA using spectrophotometry or gel electrophoresis.
Advantages:
- Scalable and cost-effective for producing large quantities of gRNA.
- Suitable for generating gRNAs for various CRISPR applications.
Disadvantages:
- Requires a DNA template with a specific promoter sequence.
- May result in gRNAs with non-canonical ends, which can affect their activity.
3.2. Chemical Synthesis
Chemical synthesis involves the direct synthesis of gRNAs using automated oligonucleotide synthesizers. This method allows for precise control over the gRNA sequence and modifications.
Procedure:
-
Design the gRNA Sequence: Determine the desired gRNA sequence, including any modifications (e.g., 2′-O-methyl modifications).
-
Order the gRNA: Order the gRNA from a commercial oligonucleotide synthesis service.
-
Receive and Resuspend the gRNA: Upon receiving the synthesized gRNA, resuspend it in a suitable buffer (e.g., RNase-free water or TE buffer).
-
Quality Control: Verify the sequence and purity of the synthesized gRNA using mass spectrometry or HPLC.
Advantages:
- High purity and sequence accuracy.
- Allows for the incorporation of modified nucleotides to enhance gRNA stability and reduce off-target effects.
- No need for a DNA template or in vitro transcription.
Disadvantages:
- More expensive than in vitro transcription, especially for long or modified gRNAs.
- Limited scalability for very large quantities.
3.3. In Vivo Expression
In vivo expression involves delivering a DNA vector encoding the gRNA into cells, where the gRNA is transcribed from the vector. This method is commonly used for stable gene editing in cell lines or in vivo applications.
Procedure:
-
Clone the gRNA Sequence: Insert the gRNA sequence downstream of a promoter (e.g., U6 or H1) in a DNA vector.
-
Transfect or Transduce Cells: Introduce the vector into cells using transfection (for cell lines) or viral transduction (for cell lines and in vivo applications).
-
Select or Screen for Positive Cells: If using a selection marker, select for cells that have integrated the vector. Alternatively, screen for cells expressing the gRNA using techniques like RT-qPCR.
Advantages:
- Suitable for stable gene editing and long-term studies.
- Allows for tissue-specific or inducible expression of the gRNA.
Disadvantages:
- Requires cloning and cell culture.
- The expression level of the gRNA may vary between cells.
3.4. Choosing the Right Method
The choice of gRNA synthesis method depends on the specific application, budget, and available resources. In vitro transcription is a cost-effective option for generating large quantities of gRNA for transient experiments. Chemical synthesis offers high purity and allows for modifications to enhance gRNA stability and reduce off-target effects. In vivo expression is suitable for stable gene editing and long-term studies.
| Method | Advantages | Disadvantages | Applications |
|---|---|---|---|
| In Vitro Transcription | Scalable, cost-effective | Requires DNA template, potential for non-canonical ends | Transient experiments, large-scale screenings |
| Chemical Synthesis | High purity, sequence accuracy, allows for modifications | More expensive, limited scalability | Applications requiring high purity, modified gRNAs |
| In Vivo Expression | Suitable for stable gene editing, allows for tissue-specific or inducible expression | Requires cloning and cell culture, variable expression levels between cells | Stable cell lines, in vivo applications |
4. Validating gRNA Activity
After synthesizing the gRNA, it is essential to validate its activity. Validation ensures that the gRNA effectively guides the Cas9 enzyme to the target site and induces the desired genomic modification. Several methods can be used to assess gRNA activity.
4.1. In Vitro Cleavage Assay
An in vitro cleavage assay is a direct method to assess the ability of the gRNA-Cas9 complex to cleave the target DNA. This assay involves incubating the gRNA-Cas9 complex with a DNA fragment containing the target sequence and then analyzing the cleavage products.
Procedure:
-
Prepare the gRNA-Cas9 Complex: Mix the synthesized gRNA with the Cas9 enzyme in a suitable buffer.
-
Prepare the Target DNA Fragment: Amplify the target DNA region by PCR or obtain it from a plasmid.
-
Incubate the Complex with the DNA Fragment: Incubate the gRNA-Cas9 complex with the target DNA fragment at 37°C for 1-4 hours.
-
Analyze Cleavage Products: Analyze the cleavage products by agarose gel electrophoresis. If the gRNA-Cas9 complex is active, the DNA fragment will be cleaved into smaller fragments.
Advantages:
- Directly assesses gRNA activity.
- Relatively quick and easy to perform.
Disadvantages:
- Does not reflect cellular conditions.
- Requires purified Cas9 enzyme and target DNA.
4.2. T7 Endonuclease I (T7E1) Assay
The T7E1 assay is a common method for detecting insertions and deletions (indels) caused by CRISPR-Cas9-mediated DNA cleavage and non-homologous end joining (NHEJ) repair. T7E1 is an enzyme that recognizes and cleaves heteroduplex DNA, which forms when wild-type and indel-containing DNA strands re-anneal.
Procedure:
-
Transfect Cells with gRNA and Cas9: Transfect cells with plasmids expressing the gRNA and Cas9.
-
Extract Genomic DNA: After 48-72 hours, extract genomic DNA from the transfected cells.
-
Amplify the Target Region: Amplify the target region by PCR using primers flanking the gRNA target site.
-
Denature and Re-anneal the PCR Product: Denature the PCR product by heating to 95°C, then allow it to re-anneal slowly to form heteroduplexes.
-
Digest with T7E1: Digest the re-annealed PCR product with T7E1 enzyme.
-
Analyze Digestion Products: Analyze the digestion products by agarose gel electrophoresis. If indels are present, T7E1 will cleave the heteroduplexes, resulting in smaller fragments.
Advantages:
- Easy to perform and relatively inexpensive.
- Can detect low levels of gene editing.
Disadvantages:
- Qualitative or semi-quantitative.
- May not detect all types of indels.
4.3. Surveyor Nuclease Assay
The Surveyor nuclease assay is similar to the T7E1 assay and is used to detect indels caused by CRISPR-Cas9. Surveyor nuclease also recognizes and cleaves heteroduplex DNA.
Procedure:
-
Transfect Cells with gRNA and Cas9: Transfect cells with plasmids expressing the gRNA and Cas9.
-
Extract Genomic DNA: Extract genomic DNA from the transfected cells.
-
Amplify the Target Region: Amplify the target region by PCR.
-
Denature and Re-anneal the PCR Product: Denature and re-anneal the PCR product to form heteroduplexes.
-
Digest with Surveyor Nuclease: Digest the re-annealed PCR product with Surveyor nuclease.
-
Analyze Digestion Products: Analyze the digestion products by agarose gel electrophoresis.
Advantages:
- Similar to the T7E1 assay.
- Can be used as an alternative if T7E1 is not available.
Disadvantages:
- Qualitative or semi-quantitative.
- May not detect all types of indels.
4.4. Sequencing
Sequencing is a quantitative method for assessing gRNA activity. This method involves amplifying the target region by PCR and then sequencing the PCR product to identify and quantify indels.
Procedure:
-
Transfect Cells with gRNA and Cas9: Transfect cells with plasmids expressing the gRNA and Cas9.
-
Extract Genomic DNA: Extract genomic DNA from the transfected cells.
-
Amplify the Target Region: Amplify the target region by PCR.
-
Prepare Sequencing Libraries: Prepare sequencing libraries from the PCR product.
-
Perform Sequencing: Perform next-generation sequencing (NGS) to analyze the sequence of the PCR product.
-
Analyze Sequencing Data: Analyze the sequencing data to identify and quantify indels. Several software tools are available for analyzing CRISPR-Cas9 sequencing data, such as CRISPResso and ICE.
Advantages:
- Quantitative and highly accurate.
- Can detect all types of indels.
- Provides detailed information about the types and frequencies of mutations.
Disadvantages:
- More expensive than other methods.
- Requires expertise in sequencing and bioinformatics analysis.
4.5. Choosing the Right Validation Method
The choice of validation method depends on the specific application and available resources. The in vitro cleavage assay is a quick and direct method for assessing gRNA activity. The T7E1 and Surveyor nuclease assays are easy and inexpensive methods for detecting indels. Sequencing is a quantitative and highly accurate method for assessing gRNA activity, but it is more expensive and requires expertise in sequencing and bioinformatics analysis.
| Method | Advantages | Disadvantages | Applications |
|---|---|---|---|
| In Vitro Cleavage Assay | Direct assessment, quick and easy | Does not reflect cellular conditions, requires purified components | Initial screening of gRNA activity |
| T7 Endonuclease I Assay | Easy to perform, inexpensive | Qualitative or semi-quantitative, may not detect all types of indels | Detecting indels in cell culture |
| Surveyor Nuclease Assay | Similar to T7E1, can be used as an alternative | Qualitative or semi-quantitative, may not detect all types of indels | Detecting indels in cell culture |
| Sequencing | Quantitative, highly accurate, detects all types of indels | More expensive, requires expertise in sequencing and bioinformatics analysis | Detailed analysis of mutations, quantifying editing efficiency |
5. Optimizing gRNA Performance
Even with careful design and validation, gRNA performance can vary. Optimizing gRNA performance involves fine-tuning experimental conditions and making adjustments to the gRNA itself to enhance its efficacy and specificity.
5.1. gRNA Modifications
Modifying gRNAs can enhance their stability, reduce off-target effects, and improve their overall performance. Common modifications include:
-
2′-O-Methyl Modifications: Introducing 2′-O-methyl groups at the 5′ and 3′ ends of the gRNA can increase its stability and resistance to degradation by cellular nucleases.
-
Phosphorothioate (PS) Modifications: Incorporating phosphorothioate linkages between nucleotides can also enhance gRNA stability.
-
Locked Nucleic Acid (LNA) Modifications: LNA modifications involve the incorporation of LNA nucleotides, which have a constrained ribose moiety. LNA modifications can increase the binding affinity of the gRNA to the target DNA and reduce off-target effects.
5.2. Adjusting Cas9 Expression Levels
The level of Cas9 expression can significantly impact the efficiency of CRISPR-Cas9-mediated gene editing. Optimizing Cas9 expression levels can improve gRNA performance.
-
Titration of Cas9 Plasmid: If using a plasmid to express Cas9, titrate the amount of plasmid transfected into cells to find the optimal dose. Too much Cas9 can lead to off-target effects, while too little Cas9 can reduce editing efficiency.
-
Using Inducible Cas9 Systems: Inducible Cas9 systems allow for controlled expression of Cas9. These systems can be used to fine-tune Cas9 expression levels and minimize off-target effects.
5.3. Optimizing Delivery Methods
The method used to deliver the gRNA and Cas9 into cells can also affect gRNA performance. Common delivery methods include:
-
Transfection: Transfection involves using chemical or physical methods to introduce the gRNA and Cas9 into cells. Common transfection reagents include Lipofectamine and electroporation.
-
Viral Transduction: Viral transduction involves using viral vectors to deliver the gRNA and Cas9 into cells. Viral vectors, such as lentiviruses and adeno-associated viruses (AAVs), can efficiently transduce a wide range of cell types.
-
Ribonucleoprotein (RNP) Delivery: RNP delivery involves pre-assembling the gRNA and Cas9 into a ribonucleoprotein complex and then delivering the complex directly into cells. RNP delivery can reduce off-target effects because the Cas9 enzyme is only active for a short period.
5.4. Optimizing Incubation Time
The incubation time after delivering the gRNA and Cas9 into cells can affect gRNA performance. Optimizing the incubation time can improve editing efficiency.
- Time Course Experiments: Perform time course experiments to determine the optimal incubation time. Extract genomic DNA at different time points after transfection or transduction and then assess gene editing efficiency using methods like T7E1 assay or sequencing.
5.5. Using Multiple gRNAs
Using multiple gRNAs targeting the same gene can increase editing efficiency. This strategy involves designing two or more gRNAs that target different sites within the same gene and then delivering all the gRNAs and Cas9 into cells.
-
Synergistic Effects: Multiple gRNAs can act synergistically to increase gene editing efficiency.
-
Improved Knockout Efficiency: Using multiple gRNAs can improve the efficiency of gene knockout by inducing larger deletions or inversions.
5.6. Considering Cellular Context
The cellular context can significantly affect gRNA performance. Factors such as cell type, chromatin structure, and DNA repair pathways can influence the efficiency of CRISPR-Cas9-mediated gene editing.
-
Cell-Type-Specific Optimization: Optimize gRNA performance for each cell type.
-
Epigenetic Modifications: Consider the epigenetic modifications at the target site.
By systematically optimizing gRNA performance, researchers can enhance the precision and efficiency of CRISPR-Cas9-mediated genome editing.
6. Ethical Considerations and Best Practices
As CRISPR technology becomes more widespread, it is essential to consider the ethical implications and adhere to best practices. Ethical considerations ensure that CRISPR technology is used responsibly and for the benefit of society.
6.1. Informed Consent
When using CRISPR technology in human cells or organisms, it is essential to obtain informed consent. Informed consent ensures that individuals understand the potential risks and benefits of the technology and have the right to make decisions about its use.
-
Transparency: Provide clear and transparent information about the purpose of the research, the methods used, and the potential outcomes.
-
Voluntary Participation: Ensure that participation is voluntary and that individuals have the right to withdraw at any time.
6.2. Data Security and Privacy
Protecting data security and privacy is essential when working with CRISPR technology. Genetic information is sensitive and must be protected from unauthorized access.
-
Secure Data Storage: Store genetic data in secure, encrypted databases.
-
Anonymization: Anonymize data whenever possible to protect individual privacy.
6.3. Responsible Use of Technology
Use CRISPR technology responsibly and for the benefit of society. Avoid using CRISPR technology for purposes that could harm individuals or society.
-
Therapeutic Applications: Focus on using CRISPR technology for therapeutic applications, such as treating genetic diseases.
-
Avoid Enhancement: Avoid using CRISPR technology for human enhancement.
6.4. Compliance with Regulations
Comply with all applicable regulations and guidelines when using CRISPR technology. Regulations vary by country and region, so it is essential to stay informed and adhere to the relevant rules.
-
Institutional Review Boards (IRBs): Seek approval from IRBs before conducting research involving CRISPR technology.
-
Governmental Oversight: Comply with governmental oversight and regulations.
6.5. Promoting Transparency and Open Communication
Promote transparency and open communication about CRISPR technology. Share research findings and engage in public discussions to increase understanding and awareness.
-
Publication of Results: Publish research findings in peer-reviewed journals.
-
Public Engagement: Engage in public discussions to increase understanding and awareness of CRISPR technology.
By adhering to ethical considerations and best practices, researchers can ensure that CRISPR technology is used responsibly and for the benefit of society. At CONDUCT.EDU.VN, we emphasize the importance of ethical conduct in scientific research and provide resources to help researchers navigate the ethical challenges associated with CRISPR technology.
7. Troubleshooting Common Issues
Even with careful planning and execution, CRISPR experiments can sometimes encounter issues. Troubleshooting common problems can help identify the cause of the issue and find a solution.
7.1. Low Editing Efficiency
Low editing efficiency is a common problem in CRISPR experiments. Several factors can contribute to low editing efficiency.
-
gRNA Design: Poor gRNA design can result in low editing efficiency. Ensure that the gRNA is designed according to the guidelines discussed earlier in this article.
-
Cas9 Expression Levels: Insufficient Cas9 expression can reduce editing efficiency. Optimize Cas9 expression levels by titrating the amount of Cas9 plasmid or using an inducible Cas9 system.
-
Delivery Method: Inefficient delivery of the gRNA and Cas9 into cells can reduce editing efficiency. Optimize the delivery method by using different transfection reagents or viral vectors.
-
Cell Type: Some cell types are more difficult to edit than others. Optimize the experimental conditions for the specific cell type being used.
7.2. High Off-Target Effects
High off-target effects can lead to unintended mutations at other locations in the genome. Several strategies can be used to reduce off-target effects.
-
gRNA Design: Design gRNAs with minimal homology to other regions of the genome. Use bioinformatics tools to scan the genome for potential off-target sites.
-
High-Fidelity Cas9 Variants: Use high-fidelity Cas9 variants that have been engineered to reduce off-target activity.
-
RNP Delivery: Use RNP delivery to reduce off-target effects.
-
Lower Cas9 Expression Levels: Lower Cas9 expression levels can reduce off-target effects.
7.3. Cell Toxicity
CRISPR-Cas9 can sometimes be toxic to cells. Several factors can contribute to cell toxicity.
-
High Cas9 Expression Levels: High Cas9 expression levels can be toxic to cells. Optimize Cas9 expression levels by titrating the amount of Cas9 plasmid or using an inducible Cas9 system.
-
Delivery Method: Some delivery methods, such as electroporation, can be toxic to cells. Use a less toxic delivery method, such as chemical transfection or viral transduction.
-
Incubation Time: Prolonged incubation times can increase cell toxicity. Optimize the incubation time to minimize cell toxicity.
7.4. No Detectable Editing
In some cases, no detectable editing may be observed. Several factors can contribute to this problem.
-
gRNA Inactivity: The gRNA may be inactive. Validate gRNA activity using an in vitro cleavage assay.
-
Cas9 Inactivity: The Cas9 enzyme may be inactive. Use a functional Cas9 enzyme.
-
Inefficient Delivery: The gRNA and Cas9 may not be efficiently delivered into cells. Optimize the delivery method.
7.5. Unexpected Mutations
In some cases, unexpected mutations may be observed. These mutations can be caused by off-target effects or other factors.
-
Off-Target Effects: Off-target effects can lead to unexpected mutations. Reduce off-target effects by designing gRNAs with minimal homology to other regions of the genome.
-
DNA Repair Pathways: DNA repair pathways can introduce unexpected mutations. Consider using inhibitors of DNA repair pathways to improve the accuracy of CRISPR-Cas9-mediated gene editing.
By systematically troubleshooting common issues, researchers can improve the success rate of their CRISPR experiments. At CONDUCT.EDU.VN, we provide resources to help researchers troubleshoot common problems and optimize their CRISPR experiments.
8. Future Trends in gRNA Design and Synthesis
The field of CRISPR technology is rapidly evolving, and new trends in gRNA design and synthesis are emerging. These trends promise to improve the efficiency, specificity, and versatility of CRISPR-Cas9-mediated genome editing.
8.1. Machine Learning for gRNA Design
Machine learning algorithms are being used to predict gRNA activity and off-target effects. These algorithms can analyze large datasets of gRNA sequences and experimental data to identify patterns and predict the performance of new gRNAs.
-
Improved Prediction Accuracy: Machine learning algorithms can improve the accuracy of gRNA design.
-
Identification of Novel gRNA Design Rules: Machine learning algorithms can identify novel gRNA design rules that are not apparent from traditional methods.
8.2. High-Throughput gRNA Screening
High-throughput gRNA screening involves testing large libraries of gRNAs to identify the most effective gRNAs for a specific target. This approach can be used to optimize gRNA design and improve editing efficiency.
-
Identification of Optimal gRNAs: High-throughput gRNA screening can identify the most effective gRNAs for a specific target.
-
Characterization of gRNA Activity: High-throughput gRNA screening can characterize the activity of large numbers of gRNAs.
8.3. Novel Cas Enzymes
Researchers are discovering and engineering novel Cas enzymes with different PAM sequences and other properties. These enzymes can expand the range of targetable sites and improve the specificity of CRISPR-Cas9-mediated genome editing.
-
Expanded Target Range: Novel Cas enzymes can expand the range of targetable sites.
-
Improved Specificity: Novel Cas enzymes can improve the specificity of CRISPR-Cas9-mediated genome editing.
8.4. RNA Modifications
RNA modifications, such as 2′-O-methyl modifications and phosphorothioate modifications, are being used to enhance gRNA stability and reduce off-target effects. New RNA modifications are being developed to further improve gRNA performance.
-
Enhanced gRNA Stability: RNA modifications can enhance gRNA stability.
-
Reduced Off-Target Effects: RNA modifications can reduce off-target effects.
8.5. Delivery Methods
New delivery methods are being developed to improve the efficiency and safety of CRISPR-Cas9 delivery. These methods include:
-
Nanoparticle Delivery: Nanoparticles can be used to deliver the gRNA and Cas9 into cells.
-
Exosome Delivery: Exosomes are small vesicles that can be used to deliver the gRNA and Cas9 into cells.
8.6. Base Editing and Prime Editing
Base editing and prime editing are new CRISPR-based technologies that allow for precise editing of DNA without double-strand breaks. These technologies offer the potential to correct genetic mutations with high precision and minimal off-target effects.
-
Precise Editing of DNA: Base editing and prime editing allow for precise editing of DNA.
-
Minimal Off-Target Effects: Base editing and prime editing have minimal off-target effects.
By staying informed about future trends in gRNA design and synthesis, researchers can continue to improve the efficiency, specificity, and versatility of CRISPR-Cas9-mediated genome editing.
9. Conclusion
Mastering how to make guide RNA for CRISPR is essential for researchers aiming to leverage this powerful tool for genome editing. By understanding the intricacies of gRNA design, synthesis, validation, and optimization, scientists can enhance the precision and efficacy of their experiments. This comprehensive guide has provided a detailed overview of the key considerations and best practices for creating effective gRNAs, ensuring ethical and responsible use of CRISPR technology.
CONDUCT.EDU.VN is committed to providing researchers with the resources and guidance needed to navigate the complexities of CRISPR technology. By staying informed about the latest advancements and adhering to ethical guidelines, we can collectively harness the potential of CRISPR to advance scientific knowledge and improve human health.
For more detailed guidance, resources, and support, visit CONDUCT.EDU.VN. Our team of experts is available to assist you with your research needs and ensure the responsible and ethical conduct of your scientific endeavors. Contact us at 100 Ethics Plaza, Guideline City, CA 90210, United States, or reach out via WhatsApp at +1 (707) 555-1234. Together, we can unlock the full potential of CRISPR technology while upholding the highest standards of ethics and integrity.
10. Frequently Asked Questions (FAQ)
1. What is a guide RNA (gRNA)?
A guide RNA (gRNA) is a short RNA sequence that directs the Cas9 enzyme to a specific location in the genome. The gRNA consists of two essential components: CRISPR RNA (crRNA) and trans-activating crRNA (tracrRNA).
2. How do I design a gRNA for CRISPR-Cas9?
To design a gRNA, select a 20-nucleotide target sequence adjacent to a PAM sequence (NGG for SpCas9). Minimize off-target effects by selecting a sequence with minimal homology to other regions of the genome.
3. What are the common methods for gRNA synthesis?
Common methods for gRNA synthesis include in vitro transcription, chemical synthesis, and in vivo expression.
4. How do I validate gRNA activity?
gRNA activity can be validated using methods such as in vitro cleavage assay, T7 Endonuclease I (T7E1) assay, Surveyor nuclease assay, and sequencing.
5. How can I optimize gRNA performance?
gRNA performance can be optimized through modifications, adjusting Cas9 expression levels, optimizing delivery methods, and considering the cellular context.
6. What are the ethical considerations when using CRISPR technology?
Ethical considerations include obtaining informed consent, protecting data security and privacy, using the technology responsibly, complying with regulations, and promoting transparency and open communication.
7. What are some common issues in CRISPR experiments and how can I troubleshoot them?
Common issues include low editing efficiency, high off-target effects, and cell toxicity. Troubleshooting involves optimizing gRNA design, adjusting Cas9 expression levels, and refining delivery methods.
8. How do machine learning algorithms aid in gRNA design?
Machine learning algorithms analyze large datasets to predict gRNA activity and off-target effects, improving the accuracy and efficiency of gRNA design.
9. What are some future trends in gRNA design and synthesis?
Future trends include machine learning for gRNA design, high-throughput gRNA screening, novel Cas enzymes, RNA modifications, and improved delivery methods.
10. Where can I find more resources and support for CRISPR technology?
Additional resources and support can be found at conduct.edu.vn, which offers comprehensive guidance, expert assistance, and ethical guidelines for CRISPR research.
