Designing guide RNAs (gRNAs) for CRISPR-Cas systems can be challenging, but it’s also essential for successful genome editing. At CONDUCT.EDU.VN, we provide comprehensive guidance on the best practices and considerations for gRNA design, ensuring accurate and efficient genome modification. Discover how to design effective gRNAs, minimize off-target effects, and optimize your CRISPR experiments with our expert insights and practical tools.
1. Understanding CRISPR and gRNA Design
The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) technology and CRISPR-associated protein (Cas) systems are revolutionizing genetic engineering. This powerful tool allows scientists to precisely edit DNA sequences within living organisms. At the heart of this technology lies the guide RNA (gRNA), a small RNA molecule that directs the Cas enzyme (typically Cas9) to a specific location in the genome. Understanding the principles of gRNA design is crucial for the success of any CRISPR-based experiment.
1.1. What is CRISPR-Cas9?
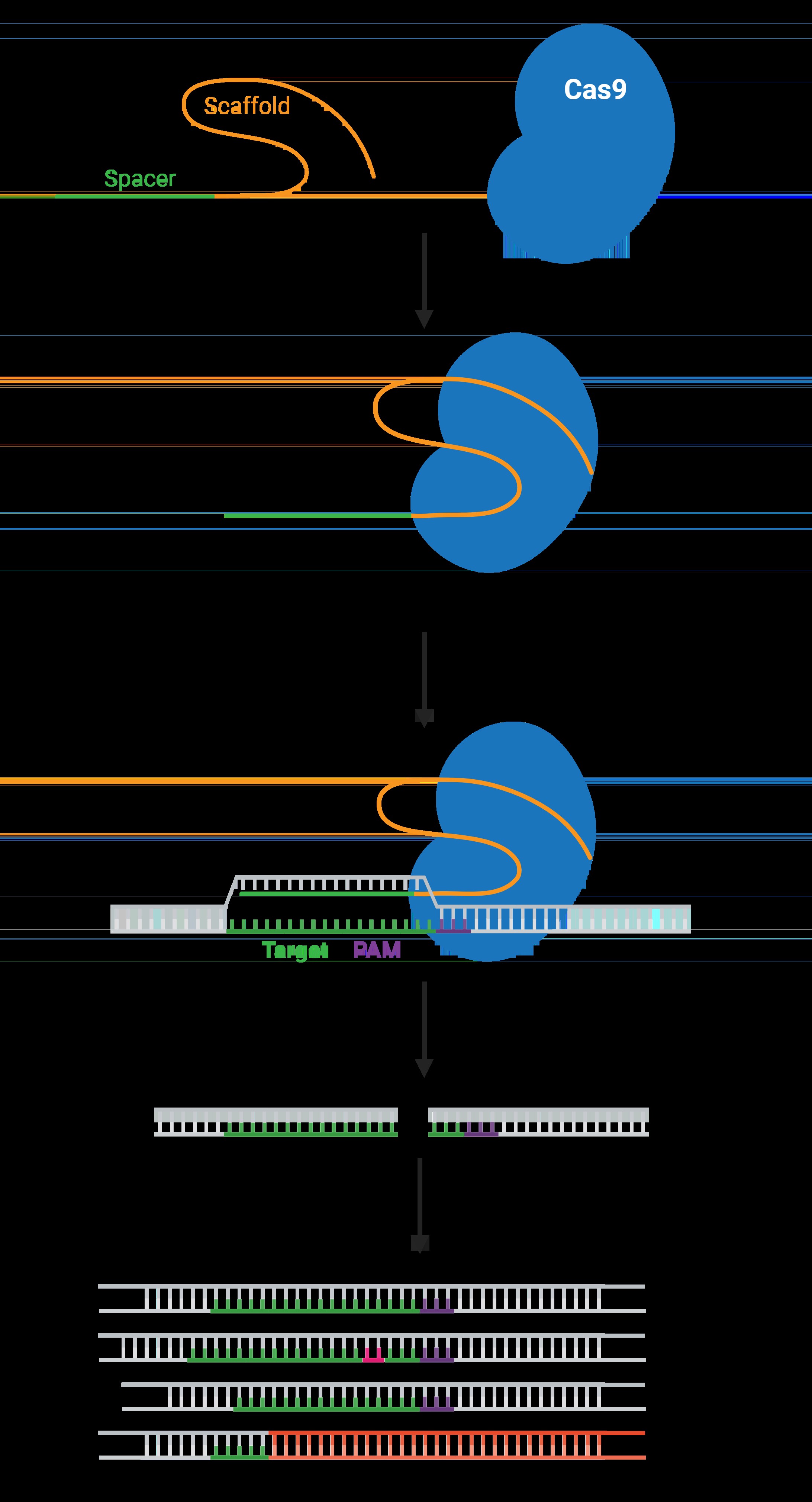
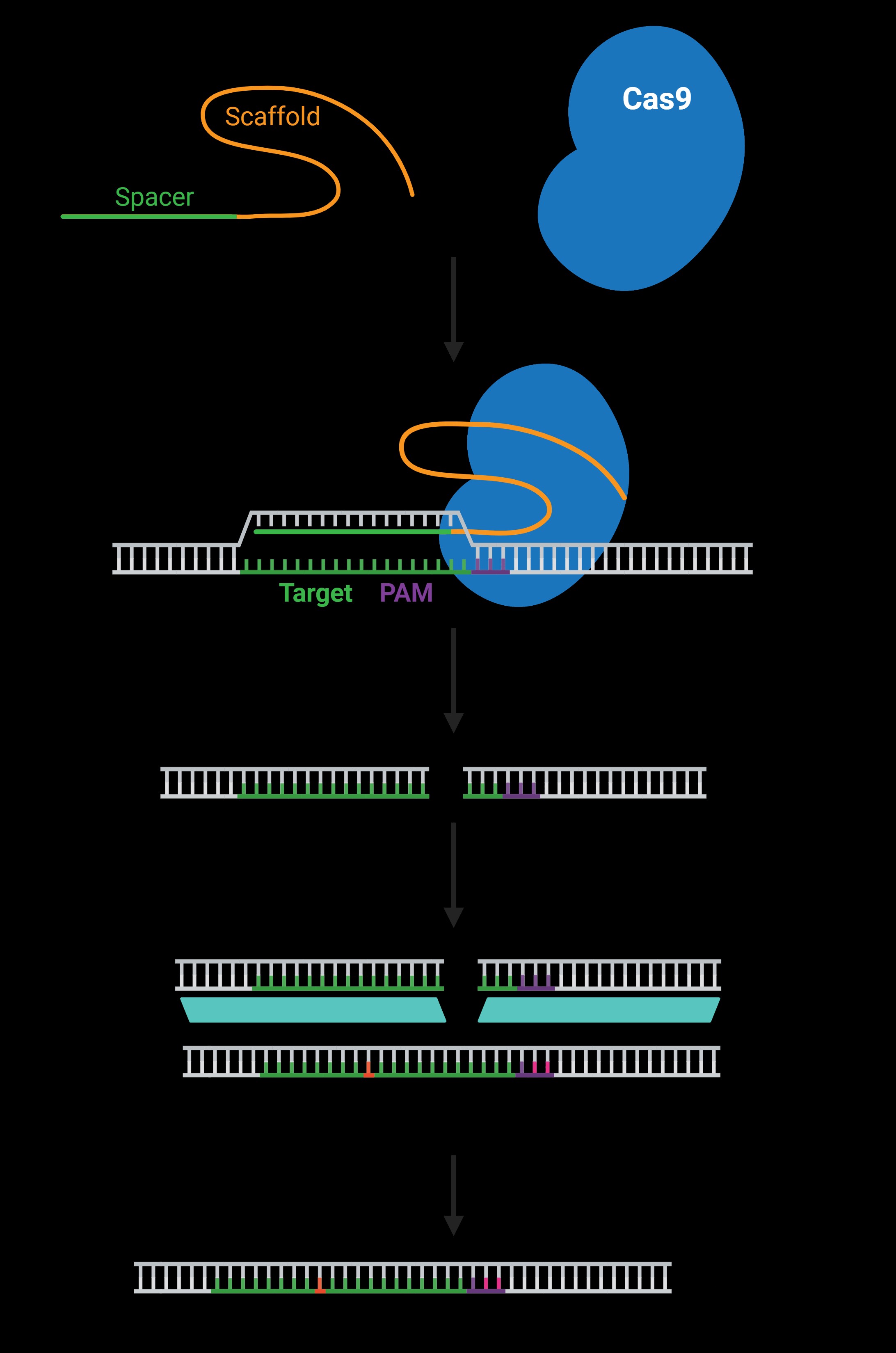
CRISPR-Cas9 is a genome editing technology derived from a natural defense mechanism used by bacteria to protect themselves from viral infections. In this system, Cas9 acts like molecular scissors, cutting DNA at a location specified by the gRNA. This targeted cutting can be used to disrupt genes, insert new DNA sequences, or correct genetic mutations. The process typically involves:
- gRNA Design: Designing a gRNA that matches the target DNA sequence.
- Complex Formation: The gRNA binds to the Cas9 protein.
- Targeting: The gRNA guides the Cas9 complex to the target DNA.
- Cleavage: Cas9 cuts the DNA at the targeted location.
- Repair: The cell’s natural repair mechanisms fix the break, often introducing mutations or incorporating a new DNA sequence.
1.2. The Role of gRNA
The gRNA is a critical component of the CRISPR-Cas9 system because it determines the specificity of the DNA editing process. A well-designed gRNA ensures that Cas9 cuts the DNA at the intended location, minimizing off-target effects. The gRNA typically consists of two parts:
- Spacer Sequence: A 20-nucleotide sequence that is complementary to the target DNA.
- Scaffold Sequence: A constant sequence that binds to the Cas9 protein.
1.3. Key Considerations for gRNA Design
Several factors must be considered when designing gRNAs to ensure effective and specific genome editing. These include:
- On-Target Activity: The gRNA should be highly active and efficiently guide Cas9 to the target site.
- Off-Target Effects: The gRNA should have minimal similarity to other regions of the genome to avoid unintended cuts.
- PAM Sequence: The protospacer adjacent motif (PAM) is a short DNA sequence required for Cas9 binding and cutting. The gRNA must be designed to target a region adjacent to the correct PAM sequence.
- Location: The location of the target site within the gene or regulatory region can affect the outcome of the edit.
- Sequence Composition: Certain sequence features can affect gRNA activity and should be considered during design.
2. Identifying Target Sites and PAM Sequences
The first step in designing a gRNA is to identify potential target sites in the genome. This involves scanning the DNA sequence for regions that are both unique and adjacent to a suitable PAM sequence. Different Cas enzymes have different PAM requirements, so it’s important to select a Cas enzyme and PAM sequence that are appropriate for your target region.
2.1. Understanding PAM Sequences
The Protospacer Adjacent Motif (PAM) is a short DNA sequence that is essential for Cas9 binding and cleavage. The most commonly used Cas9 enzyme, Streptococcus pyogenes Cas9 (SpCas9), recognizes the PAM sequence 5′-NGG-3′, where N can be any nucleotide. Other Cas enzymes have different PAM requirements, which can expand the range of targetable sites.
| Cas Enzyme | PAM Sequence | Source | Advantages | Disadvantages |
|---|---|---|---|---|
| SpCas9 | 5′-NGG-3′ | Streptococcus pyogenes | Widely used, well-characterized | Limited by NGG PAM |
| SaCas9 | 5′-NNGRRT-3′ | Staphylococcus aureus | Smaller size, different PAM expands targeting range | Lower activity compared to SpCas9 |
| NmeCas9 | 5′-RCCGAC-3′ | Neisseria meningitidis | Unique PAM, expands targeting range | Less efficient than SpCas9, more off-target effects |
| Cas12a (Cpf1) | 5′-TTTN-3′ | Acidaminococcus and Lachnospiraceae | Creates sticky ends, useful for precise insertions | Requires different design considerations |
| Engineered Cas9s | Varies | Various (engineered variants) | Tailored PAM specificities, improved on-target activity, reduced off-target effects, expanded targeting | May require specific experimental conditions, potential for unforeseen effects, continuous optimization |
2.2. Strategies for Identifying Target Sites
Identifying suitable target sites involves using bioinformatics tools and databases to scan the genome for sequences that meet the following criteria:
- Uniqueness: The 20-nucleotide spacer sequence should be unique to the target gene or region to avoid off-target effects.
- PAM Proximity: The target site must be immediately adjacent to a suitable PAM sequence for the chosen Cas enzyme.
- Location: The location of the target site within the gene or regulatory region should be chosen based on the desired outcome (e.g., disrupting the coding sequence, targeting a promoter region).
- Accessibility: The target site should be accessible to the Cas9 complex, considering factors like chromatin structure and DNA methylation.
2.3. Tools and Databases for Target Site Selection
Several online tools and databases can assist in identifying potential target sites. These tools typically allow you to input a DNA sequence and will output a list of potential gRNAs, along with information on their predicted activity and off-target potential. Some popular tools include:
- CRISPR Design Tool (MIT): A comprehensive tool for designing gRNAs for various Cas enzymes.
- CHOPCHOP: A user-friendly tool that provides information on gRNA activity, specificity, and off-target potential.
- Benchling: A molecular biology software platform that includes CRISPR design tools and sequence analysis capabilities.
- CRISPRz: A tool providing comprehensive gRNA design, off-target analysis, and batch processing capabilities.
These tools often incorporate algorithms that predict gRNA activity and off-target effects, helping you to prioritize the best candidates for your experiment.
3. Designing gRNAs for Specific Applications
The specific requirements for gRNA design can vary depending on the application, such as gene knockout, gene editing, or transcriptional modulation. Each application has unique considerations that must be taken into account to achieve the desired outcome.
3.1. gRNA Design for Gene Knockout
Gene knockout involves disrupting a gene’s function by introducing insertions or deletions (indels) at the target site. To maximize the chances of a successful knockout, consider the following:
- Target Coding Region: Target the coding region of the gene to disrupt protein production.
- Avoid N-Terminal Targets: Avoid targeting sites near the N-terminus of the protein to prevent the cell from using an alternative start codon.
- Avoid C-Terminal Targets: Avoid targeting sites near the C-terminus of the protein to maximize the chances of creating a non-functional allele.
- Multiple gRNAs: Design multiple gRNAs targeting different locations within the gene to increase the likelihood of a successful knockout.
3.2. gRNA Design for Gene Editing (HDR)
Gene editing involves introducing specific changes to the DNA sequence, such as inserting a fluorescent tag or correcting a mutation. This is typically achieved through homology-directed repair (HDR), which requires an exogenous DNA template. The efficiency of HDR decreases dramatically as the distance between the cut site and the desired edit increases.
- Proximity to Edit: Target sites within ~30 nucleotides of the desired edit to maximize HDR efficiency.
- PAM Options: Consider using alternative Cas enzymes with different PAM sequences to expand targeting options.
- Exogenous Template: Design an exogenous DNA template with homology arms flanking the desired edit and a PAM mutation to prevent recutting.
3.3. gRNA Design for Base Editing and Prime Editing
Base editing and prime editing are newer technologies that offer alternatives to HDR for introducing specific edits. Base editors convert one base pair to another without creating double-strand breaks, while prime editors can insert or delete DNA sequences at the target site.
- Base Editing: The target base must be within a 5-10 nucleotide window relative to the PAM sequence. Be aware of potential bystander edits if there are other target bases in the window.
- Prime Editing: Requires a nearby PAM sequence, and the user may need to optimize various parameters to achieve the desired edit.
3.4. gRNA Design for CRISPRa and CRISPRi
CRISPR activation (CRISPRa) and CRISPR interference (CRISPRi) are used to modulate gene expression at the transcriptional level. A nuclease-dead Cas9 (dCas9) is directed to the promoter region of a target gene, where it can either activate or repress transcription.
- CRISPRa: Target a ~100-nucleotide window upstream of the transcription start site (TSS).
- CRISPRi: Target a ~100-nucleotide window downstream of the TSS.
- Accurate TSS Information: Use accurate TSS information from databases like FANTOM to ensure proper targeting.
4. Minimizing Off-Target Effects
Off-target effects occur when the gRNA directs Cas9 to cut DNA at unintended locations in the genome. This can lead to undesirable mutations and potentially harmful consequences. Minimizing off-target effects is a critical aspect of gRNA design.
4.1. Strategies for Reducing Off-Target Activity
- Unique Target Sequences: Choose target sequences that are unique to the intended gene or region, with minimal similarity to other parts of the genome.
- Off-Target Analysis Tools: Use off-target analysis tools to identify potential off-target sites and assess their similarity to the target site.
- High-Fidelity Cas Enzymes: Consider using high-fidelity Cas enzymes, which have been engineered to reduce off-target activity.
- Paired Nickases: Use paired Cas9 nickases, which require two gRNAs to create a double-strand break, reducing the likelihood of off-target cutting.
- Titration: Optimize the concentration of Cas9 and gRNA to minimize off-target effects while maintaining on-target activity.
4.2. Using Bioinformatics Tools for Off-Target Prediction
Several bioinformatics tools can help predict potential off-target sites. These tools use algorithms to scan the genome for sequences that are similar to the gRNA target sequence and predict the likelihood of Cas9 cutting at those sites. Some popular tools include:
- CRISPR Design Tool (MIT)
- CHOPCHOP
- Cas-OFFinder
- Benchling
By using these tools, you can identify and avoid gRNAs with a high potential for off-target activity.
4.3. Validating gRNA Specificity
Even with careful design and off-target prediction, it’s important to validate the specificity of the gRNA experimentally. This can be done using methods such as:
- Targeted Deep Sequencing: Sequence the predicted off-target sites to check for unintended mutations.
- Whole-Genome Sequencing: Perform whole-genome sequencing to identify any off-target mutations across the entire genome.
- Cellular Assays: Use cellular assays to assess the functional consequences of off-target effects.
5. Optimizing gRNA Efficacy
In addition to minimizing off-target effects, it’s important to optimize gRNA efficacy to ensure efficient on-target activity. Several factors can influence gRNA efficacy, including sequence composition, secondary structure, and chromatin accessibility.
5.1. Factors Affecting gRNA Efficacy
- GC Content: gRNAs with a GC content between 30% and 70% tend to be more active.
- Secondary Structure: Avoid gRNAs with strong secondary structures, as these can interfere with Cas9 binding.
- Position within Gene: Target sites in exons located closer to the 5′ end of the gene tend to be more effective.
- Chromatin Accessibility: Target sites in regions of open chromatin are more accessible to Cas9 and tend to be more active.
5.2. Predicting gRNA Activity
Several algorithms and models have been developed to predict gRNA activity based on sequence features. These models can help prioritize gRNAs for experimental testing. Some popular prediction tools include:
- Doench Score (Rule Set 2): A widely used model that incorporates sequence features and position within the gene to predict gRNA activity.
- CRISPRscan: A model that uses a convolutional neural network to predict gRNA activity.
- DeepCRISPR: A deep learning-based model that predicts gRNA activity based on sequence features and chromatin accessibility.
5.3. Validating gRNA Activity
The best way to validate gRNA activity is through experimental testing. This can be done using methods such as:
- T7 Endonuclease I Assay: A simple assay to detect indels at the target site.
- Mismatch Cleavage Assay: A more sensitive assay to detect indels.
- Next-Generation Sequencing: Sequence the target region to quantify the frequency of indels or other edits.
- Functional Assays: Use functional assays to assess the effect of the edit on gene expression or cellular phenotype.
6. Tools and Resources for gRNA Design
Numerous tools and resources are available to assist with gRNA design, off-target prediction, and activity prediction. These tools can streamline the design process and help you select the best gRNAs for your experiment.
6.1. Online gRNA Design Tools
- CRISPR Design Tool (MIT): A comprehensive tool for designing gRNAs for various Cas enzymes.
- Features: On-target scoring, off-target analysis, support for multiple Cas enzymes.
- Benefits: Widely used, comprehensive, reliable.
- CHOPCHOP: A user-friendly tool that provides information on gRNA activity, specificity, and off-target potential.
- Features: Easy to use, integrates multiple prediction algorithms, provides detailed information on potential off-target sites.
- Benefits: User-friendly, comprehensive, reliable.
- Benchling: A molecular biology software platform that includes CRISPR design tools and sequence analysis capabilities.
- Features: Integrated platform, includes sequence analysis, cloning, and CRISPR design tools.
- Benefits: Comprehensive, integrated, collaborative.
- CRISPRz: A tool providing comprehensive gRNA design, off-target analysis, and batch processing capabilities.
- Features: Comprehensive gRNA design, off-target analysis, and batch processing capabilities.
- Benefits: Comprehensive, efficient, versatile.
6.2. Databases for Target Site Selection
- Ensembl: A comprehensive database of genomic information, including gene annotations, transcript sequences, and regulatory elements.
- UCSC Genome Browser: A powerful tool for visualizing genomic data, including gene annotations, ChIP-seq data, and other functional genomics data.
- FANTOM: A database of transcription start sites (TSSs) based on CAGE-seq data.
6.3. Software for Sequence Analysis
- SnapGene: A molecular biology software that includes sequence analysis, cloning, and primer design tools.
- Geneious Prime: A comprehensive software for molecular biology and sequence analysis.
7. Case Studies and Examples
To illustrate the principles of gRNA design, let’s look at a few case studies and examples.
7.1. Case Study 1: Knockout of the TP53 Gene in Human Cancer Cells
The TP53 gene is a tumor suppressor gene that is frequently mutated in human cancers. To study the effects of TP53 knockout in cancer cells, researchers designed gRNAs targeting the coding region of the TP53 gene.
- Target Site Selection: gRNAs were designed to target exons 4, 5, 7, and 8 of the TP53 gene.
- Off-Target Analysis: Off-target analysis was performed using the CRISPR Design Tool to minimize the risk of off-target effects.
- Validation: The activity of the gRNAs was validated using the T7 Endonuclease I assay and sequencing.
- Results: Knockout of TP53 resulted in increased cell proliferation and resistance to chemotherapy.
7.2. Example: Editing the CCR5 Gene in Human T Cells
The CCR5 gene encodes a chemokine receptor that is used by HIV to enter T cells. Editing the CCR5 gene can confer resistance to HIV infection.
- Target Site Selection: A gRNA was designed to target the CCR5 gene near the start codon.
- HDR Template: An HDR template was designed to introduce a frameshift mutation in the CCR5 gene.
- Delivery: The gRNA, Cas9, and HDR template were delivered to human T cells using viral vectors.
- Results: Editing of the CCR5 gene resulted in resistance to HIV infection.
8. Frequently Asked Questions (FAQ)
1. What is the optimal length of a gRNA spacer sequence?
The most common length is 20 nucleotides, which provides a good balance between specificity and activity.
2. How close should the target site be to the PAM sequence?
The target site should be immediately adjacent to the PAM sequence.
3. How can I minimize off-target effects?
Use unique target sequences, perform off-target analysis, and consider using high-fidelity Cas enzymes.
4. What is the ideal GC content for a gRNA?
A GC content between 30% and 70% is generally recommended.
5. How can I predict gRNA activity?
Use prediction algorithms such as the Doench score or CRISPRscan.
6. How can I validate gRNA activity?
Use assays such as the T7 Endonuclease I assay or next-generation sequencing.
7. Can I use multiple gRNAs to target the same gene?
Yes, using multiple gRNAs can increase the likelihood of a successful knockout or edit.
8. What is the role of the PAM sequence?
The PAM sequence is required for Cas9 binding and cleavage.
9. What are some common off-target analysis tools?
CRISPR Design Tool, CHOPCHOP, and Cas-OFFinder.
10. How do I choose between different Cas enzymes?
Consider the PAM requirements, size, and activity of the Cas enzyme.
9. Conclusion
Designing effective gRNAs for CRISPR-Cas systems is crucial for successful genome editing. By considering factors such as target site selection, PAM sequences, off-target effects, and gRNA efficacy, you can optimize your experiments and achieve the desired outcomes. At CONDUCT.EDU.VN, we are committed to providing you with the knowledge and resources you need to master CRISPR technology.
Remember to carefully design your gRNAs, validate their activity and specificity, and interpret your results with caution. With the right tools and techniques, you can harness the power of CRISPR to make groundbreaking discoveries in biology and medicine.
For more detailed information and guidance on gRNA design, visit CONDUCT.EDU.VN. Our resources include detailed articles, tutorials, and expert advice to help you navigate the complexities of CRISPR technology.
Contact Information:
- Address: 100 Ethics Plaza, Guideline City, CA 90210, United States
- WhatsApp: +1 (707) 555-1234
- Website: CONDUCT.EDU.VN
By following these guidelines and leveraging the resources available at conduct.edu.vn, you can confidently design gRNAs for your CRISPR experiments and contribute to the advancement of genetic research.